3.1
Genetic Algorithm
The genetic algorithm processing component is a part of every child process. It uses its processing abilities to improve its actual population. Its input consists of three supportive objects (see Fig. 3.1-1):
-
Parameter File is a memory representation of the parameters of the genetic algorithm. It provides an access to these parameters during the computation. The parameters are read from the (disc) parameter file, previously created by the Parameter Editor component (for details see section 3.3).
-
Population is a set of genes, the genetic algorithm should work on. These genes are intended as a first actual population used for the computation. If no initial population is specified at the start-up, a new one is created and initialized randomly.
-
Fitness Function is an object providing the fitness evaluation (the quality determination) of a gene. This object knows also the format and encoding of genes.
The genetic algorithm processing component applies iteratively a sequence of generalized genetic operators to the population. We call them GAtools (see paragraph GAtools). The sequence can vary with respect to parameters i.e. the user can define the sequence. This sequence is repeated until the Control Mechanisms stops the computation. The Control Mechanisms include the user interaction, the termination conditions and a basic error capturing. During and after the computation the statistical data are collected, so the user can monitor the computation. The result of the genetic algorithm is the population. This is either the population that has satisfied the termination conditions or the actual population when the computation was stopped for another reason. This result population is stored to a special parameter file.
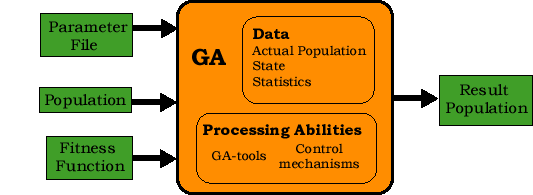
Figure 3.1-1: The scheme of the GA-processing component of the system.
3.1.1
GAtools
GAtools are generalized genetic operators. They are designed for changing
the population and for improving its overall quality. They modify the sets
of genes or the genes themselves. The order of the GAtools used during
the computation of the genetic algorithm strongly influences the performance of the algorithm.
We divide the GAtools into three basic groups: unoms, functions and selectors.
Unoms
These GAtools make changes inside a single population. Thus just one population
is required as an argument. So far we have implemented the following unoms:
-
Empty deletes all genes from the population.
-
Sizer changes the number of genes inside the population. It helps
to hold the size of the population as required by the genetic algorithm.
This is achieved by cloning or deleting the existing genes in the population.
-
Elitism inserts previously stored genes to the current population.
Then it stores the best genes from this population for later use.
Functions
These GAtools require two populations as arguments. The first one is intended
as incoming, the second one is intended as outcoming. A function first
takes the given number of genes from the incoming population. Then it performs
certain changes (or do not) with these genes. At the end it places the
result into the outcoming population. Actually, it is possible to change
the genes in the outcoming population too. We have already implemented
these functions:
-
Crossover combines two genes into two new ones. Both genes are
divided into two parts at the same randomly chosen crossing position. By
swapping the proper parts between the genes there are created two new genes.
-
Mutation randomly change some genes.
-
Pump is like the unom Sizer, but the missing genes are taken from
the incoming population.
-
Copy copies all the genes from the incoming to the outcoming population. The original genes remain members of the incoming population and their exact copies are inserted into the outcoming population.
-
Move moves all the genes from the incoming to the outcoming population. The original genes are inserted into the outcoming population. The size of the incoming population is reduced to zero.
In addition we support a special subgroup of functions called GAtables.
These are, in fact, sequences of GAtools. The user can define this sequence
in a parameter file. This allows the user substantially change the computation
of the genetic algorithm. The execution of these GAtables means a call
of each GAtool sequentially in the specified order. This mechanism enables to change the genetic algorithm without recompiling the system kernel. So far we have implemented
these GAtables:
-
Single represents a simple sequence of GAtools. This function
is intended to be a basic genetic operator, used in every regular genetic
algorithm. Any change in the sequence of GAtools, enabling more complex
computation, is possible only within this simple GAtable.
-
Cycle iteratively executes all the GAtools in its sequence.
-
Random Fork consists of two sequences of GAtools, where only one
of them is randomly chosen to proceed.
-
If Fork also consists of two sequences of GAtools and a condition.
Which one is going to be used is decided according to the condition.
-
Count Fork consists of two sequences of GAtools and one natural
number n. For a n calls of this GAtable the first sequence is executed
and then the second one.
-
Alternate as previous, but the second sequence has also specified
a number of executions and then the first sequence is executed again.
Selectors
These GAtools select a specified number of genes from incoming population.
They copy or move the selected genes into the outcoming population. Selectors
require four arguments: two populations, the number of genes to be selected
and the flag specifying whether to copy or to move them. The difference between
the implemented selectors is the strategy applied for the selection. So
far we have implemented these selectors:
-
Random selects the genes randomly. It does not consider their
properties.
-
Bests selects the fittest genes.
-
Worsts selects the least fit genes.
-
Weighted Random selects the genes with the respect to their fitness
value. The higher fitness value of a gene means the higher probability
of its selection (roulette wheel).
-
Inverted Weighted Random same as the previous, but with inverted
relation i.e. the higher fitness value of a gene means the lower probability
of its selection.

